
CRISPR-based gene editing just got safer with the design of a new SuperFi-Cas9 variant that more readily discriminates against mismatches and retains the accuracy and speed of naturally occurring Cas9.
A recent study led by Jack Bravo and his team at The University of Texas at Austin (TX, USA) reveals the redesign of Cas9, a key component of the CRISPR-based gene-editing tool, to make the technology safer. By eliminating the risks involved in CRISPR/Cas9 gene editing and making improvements by recognizing mismatches, the team have provided a proof of concept for the design of next-generation Cas9 variants – without compromising rates of rapid on-target DNA cleavage.
CRISPR/Cas9 gene editing carries some risk as the molecular machinery sometimes changes the wrong section of the host genome, which introduces the possibility that when attempting to repair this genetic mutation, the process may create a new mutation in another section. Bravo’s team has redesigned the Cas9 component to overcome this challenge, creating the new SuperFi-Cas9.
The new SuperFi-Cas9 variant is 4000–times less likely to cut off-target sites on DNA whilst functioning as rapidly as naturally–occurring Cas9 – a factor that has been compromised in previous attempts to create a new Cas9 variant with better discrimination against mismatches.
International Women’s Day with Joyce Van Eck
Joyce Van Eck is a researcher at the Boyce Thompson Institute and discusses how CRISPR/Cas9 can be used in model plants like the tomato to study gene function and how this relates to phenotypes.
Bravo compares the new SuperFi-Cas9 variant to previous attempts using a self-driving car analogy: most models are safe but have a limited top speed. Likewise, other attempts to engineer an improved Cas9 have sacrificed speed, but not SuperFi-Cas9.
“SuperFi-Cas9 is like a self-driving car that has been engineered to be extremely safe, but it can still go at full speed,” Bravo explained.
CRIPSR-based gene-editing tools are adapted from naturally arising systems in bacteria. Nature is imperfect, and so mismatches can occur when the Cas9 protein incorrectly cuts a section of DNA with a similar amino acid pattern to the target sequence. A mistake like this can have severe consequences in gene editing.
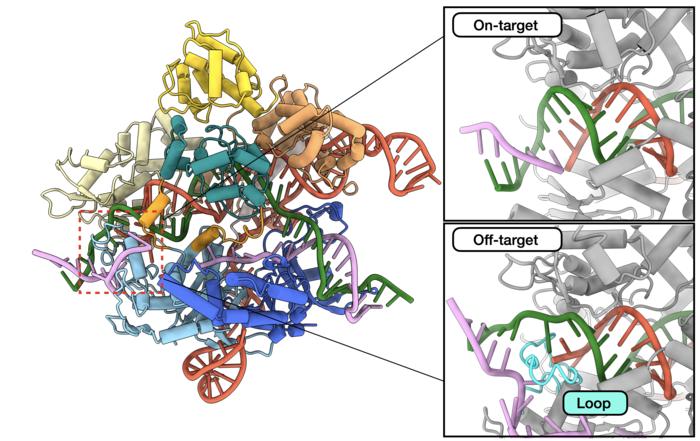
The researchers employed a technique called kinetics-guided structure determination. Using cryo-EM, the team took snapshots of the Cas9 protein in action as it encountered mismatched DNA. In doing so, the team found that when Cas9 encounters a mismatch, it has a finger-like structure that can sometimes step in and stabilize the DNA, making it appear as though it were the correct sequence, even though it’s not.
“This was something that I could never have, in a million years, imagined in my mind would have happened,” co-senior author, David Taylor, said.
The team then redesigned this finger-like structure on Cas9 to push away from the DNA, rather than stabilizing it, to prevent Cas9 from continuing the cutting and editing of the mismatch sequence. Not only does the SuperFi-Cas9 variant cut the right target as effectively and as fast as naturally– occurring Cas9 but is far less likely to cut the wrong target. The team is now collaborating with other researchers to test the technology for gene editing in living cells.
“This really could be a game changer in terms of a wider application of the CRISPR Cas systems in gene editing,” reported co-senior author of the study Kenneth Johnson.